Team SJTU-Software 2025 Official Software Tool
This repository contains the complete source code for CASPIA, an AI-powered platform designed to revolutionize synthetic biology research through intelligent automation, knowledge retrieval, and metabolic modeling.
Team Wiki: Visit our iGEM Wiki
- Overview
- Key Features
- Architecture
- Installation
- Usage
- Modules Description
- Examples
- Project Structure
- Contributing
- Testing
- Citation
- Authors and Acknowledgments
- License
- Contact
CASPIA (Cell-Automated Synthetic Pathway Intelligent Architecture) is an integrated AI-native software platform developed by Team SJTU-Software for the iGEM 2025 competition. The platform establishes a computational foundation for digital cell twins, unifying automated genome-scale modeling, high-precision parameter prediction, intelligent agent orchestration, and vision-enhanced literature retrieval.
CASPIA enables researchers to move beyond fragmented trial-and-error workflows by providing:
- GEMFactory: An automated pipeline that transforms raw genomes into parameter-enriched genome-scale metabolic models (ecGEMs/etcGEMs), incorporating kinetic and thermodynamic parameters such as kcat and Topt.
- CASPred: A multimodal predictive engine that integrates protein sequence and structure representations to complete missing kinetic parameters with quantified uncertainty.
- CASPIAgent: A natural-language-driven AI agent that plans and executes complex toolchains for gene annotation, model construction, parameter completion, and strain design optimization.
- CASPIA-RAG: A vision-augmented Retrieval-Augmented Generation system capable of analyzing both text and figures from scientific literature to provide accurate, evidence-grounded answers.
Conventional synthetic biology workflows face major limitations:
- Fragmented toolchains with inconsistent interfaces
- Manual, error-prone curation of missing kinetic parameters
- Difficulty in integrating knowledge hidden in figures, tables, and large literature corpora
- High technical barriers for non-expert users
CASPIA addresses these challenges by delivering a unified and intelligent framework that:
- ✅ Automates genome-to-model pipelines with standardized interfaces
- ✅ Completes missing parameters using cutting-edge predictive models
- ✅ Provides intuitive natural language interaction through an AI agent
- ✅ Preserves and interprets visual data from scientific publications
- ✅ Supports reproducible, traceable, and scalable metabolic engineering workflows
By compressing the Design-Build-Test-Learn (DBTL) cycle into an end-to-end digital workflow, CASPIA empowers researchers to achieve predictive, high-precision strain design and accelerates the realization of digital cell twins in synthetic biology.
- Natural language interface for complex synthetic biology tasks
- Automated orchestration of toolchains for gene annotation, GEM construction, parameter completion, and strain design
- Task planning → execution → verification workflow with exception rollback
- Context-aware reporting with full traceability of inputs, outputs, and data sources
- End-to-end automated pipeline: raw genome → parameterized GEM (ecGEM/etcGEM)
- Integration of gene annotation (GeneMarkS), protein alignment (Diamond), and metabolic network reconstruction (CarveMe)
- Automated parameter injection through database retrieval (BRENDA, KEGG, BiGG) and CASPred predictions
- Multi-scale optimization:
- Gene-level strategies (FBA, FSEOF, OptKnock)
- Protein-level mutation design (Deep Mutational Scanning with MoE PLMs: ProSST, ESM2, ProtSSN, SaProt)
- Standardized outputs in SBML and traceable reports for reproducibility
- High-precision predictive engine for missing kinetic and thermodynamic parameters (kcat, Topt)
- Multimodal architecture combining protein sequence embeddings (ESMC-300M) and structural features (GVP)
- Cross-attention fusion of sequence and structure for accurate enzyme–substrate interaction modeling
- Ensemble learning with uncertainty quantification, providing both predicted values and confidence intervals
- Continuously improved by incorporating new wet-lab data into training sets
- Vision-enhanced Retrieval-Augmented Generation for scientific literature
- PDF → Markdown structured parsing with figure/table extraction
- Image-to-text semantic captioning using vision models
- Context-preserving segmentation and embedding into Chroma vector database
- Expert Mode with cross-attention re-ranking for precise, evidence-grounded retrieval
- Accurate, cited answers integrating both textual and visual evidence
- Real-time monitoring of CASPIA computational workflows
- Visualization of job progress, status, and error recovery
- Centralized log collection for reproducibility and debugging
- Result aggregation and export for downstream analysis
System Requirements:
- Operating System: Linux (Ubuntu 20.04+), macOS (10.15+), or WSL2 on Windows
- Storage: 20GB+ free space
- RAM: 16GB minimum (32GB recommended)
- GPU: CUDA-compatible GPU with 16GB VRAM minimum (NVIDIA 4090 recommended)
Software Dependencies:
- Python: 3.10
- CUDA Toolkit 12.x (12.8 recommended)
First, install Miniconda by following the official guide: Installing Miniconda - Anaconda. Then create and configure a virtual environment. The following steps use a Linux system as an example:
# Using conda (recommended)
conda create -n caspia python=3.10
conda activate caspia
# Or using venv
python3 -m venv venv
source venv/bin/activate # On Windows: venv\Scripts\activate
# Install PyTorch with CUDA support (adjust CUDA version as needed)
pip install torch==2.7.1 torchvision==0.22.1 --index-url https://download.pytorch.org/whl/cu128
pip install torch-scatter -f https://data.pyg.org/whl/torch-2.7.1+cu128.html
pip install torch-cluster -f https://data.pyg.org/whl/torch-2.7.1+cu128.html
pip install torch-geometric
# diamond
conda install -c bioconda -c conda-forge diamond=2.1.13
# git clone
git clone git@gitlab.igem.org:2025/software-tools/sjtu-software.git
cd sjtu-software
# Install project dependencies
pip install -r requirements.txt
Note: For CUDA version compatibility, download the matching PyTorch version from the official archive: Previous PyTorch Versions.
- Visit https://huggingface.co/victorzhu30/CASPred/tree/main.
- Download the model folder to your local machine.
- Move it into the src/CASPred/ directory.
huggingface-cli download victorzhu30/CASPred
Download the Java JDK compatible with your OS from the official site: Java Downloads | Oracle. The following steps use JDK 25 as an example:
# 1. Navigate to your home directory
cd ~
# 2. Download the JDK 25 archive
wget https://download.oracle.com/java/25/latest/jdk-25_linux-x64_bin.tar.gz
# 3. Extract the archive
tar -zxvf jdk-25_linux-x64_bin.tar.gz
# 4. Edit the .bashrc file to configure environment variables
vim ~/.bashrc
# 5. Add the following lines to .bashrc (set JAVA_HOME to your JDK path)
export JAVA_HOME=~/jdk-25
export PATH=$JAVA_HOME/bin:$PATH
# 6. Save and exit Vim: Press "Esc" → type ":wq" → press "Enter"
# 7. Apply the configuration changes
source ~/.bashrcCheck if Java is installed correctly by running:
java -versionA successful installation will return output similar to:
java 25 2025-09-16 LTS
Java(TM) SE Runtime Environment (build 25+37-LTS-3491)
Java HotSpot(TM) 64-Bit Server VM (build 25+37-LTS-3491, mixed mode, sharing)GeneMarkS requires an official download link (obtained by filling out a form on the website). Follow these steps:
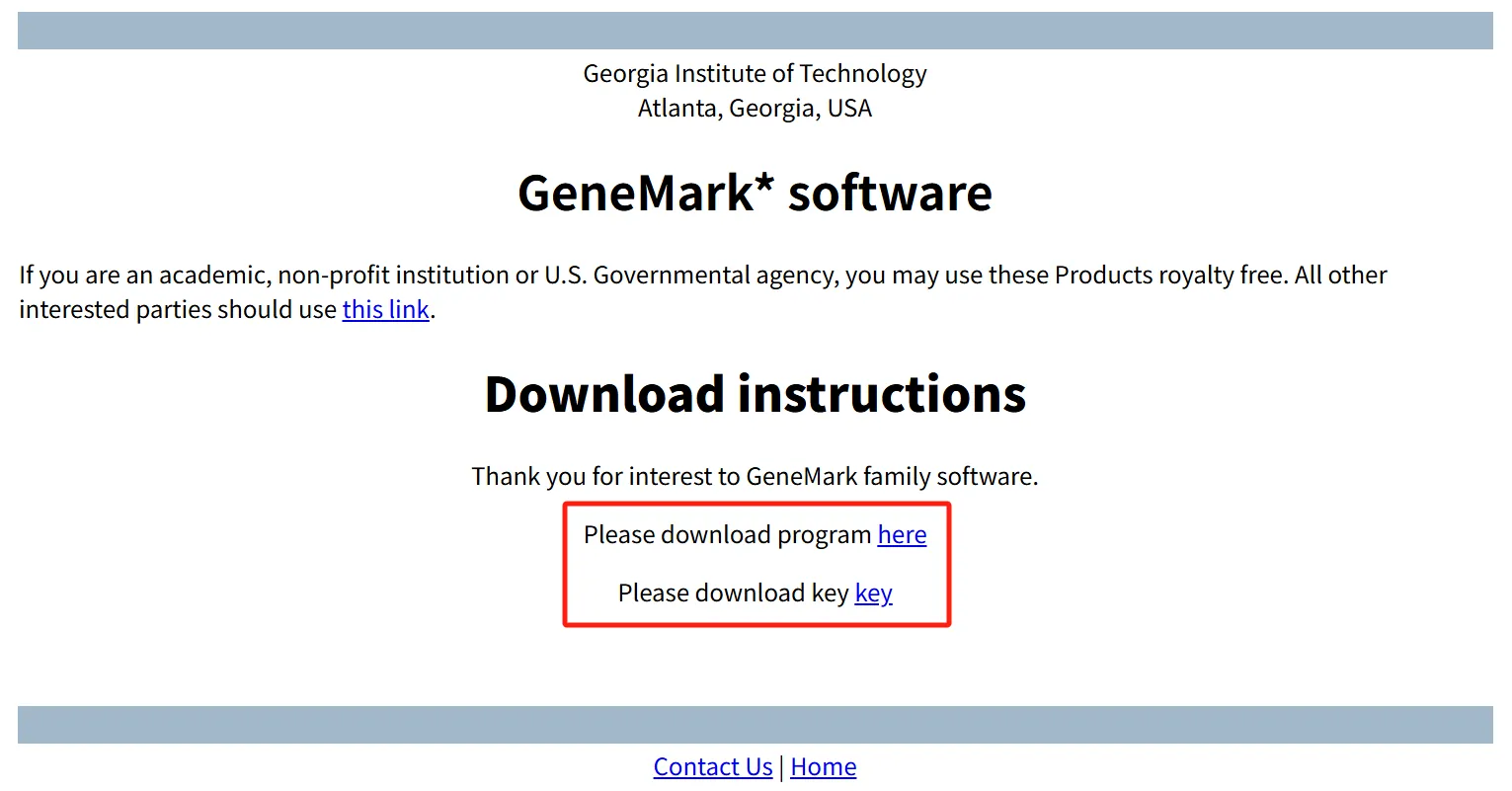
- Visit the GeneMark download page: GeneMark™ download.
- Select GeneMarkS-2 version 1.15_1.25_lic (choose the OS matching your system).
- Fill in the required personal information at the bottom of the page to generate download links.
- Right-click the links for the software package and license key, then select "Copy link address" to download them to your server.
# 1. Navigate to your home directory
cd ~
# 2. Download the GeneMarkS package and license key (replace links with your copied ones)
wget http://genemark.bme.gatech.edu/tmp/GMtool_6ghoQ/gms2_linux_64.tar.gz
wget http://genemark.bme.gatech.edu/tmp/GMtool_6ghoQ/gm_key.gz
# 3. Extract the software package
tar -zxvf gms2_linux_64.tar.gz
# 4. Install the license key (required for GeneMarkS to run)
gunzip -c gm_key.gz > ~/.gmhmmp2_key
# 5. Add GeneMarkS to system PATH (edit .bashrc)
vim ~/.bashrc
# 6. Add the following line to .bashrc (update path if needed)
PATH=$PATH:~/gms2_linux_64
# 7. Apply the PATH configuration
source ~/.bashrcEnsure the target directory has at least 2GB of free space (the model size is ~1.33GB). Follow these steps:
# Set the Hugging Face cache directory (store model here; update path as needed)
export HF_HOME=/home/shenmaa/huggingface_cache- Create a Python script named `download_esmc_300m.py` with the following content:
from esm.models.esmc import ESMC
# Download and load the ESMC-300M model (saved to HF_HOME)
model = ESMC.from_pretrained("esmc_300m")-
Run the script to start the download:
python download_esmc_300m.py
Download Notes: The model size is 1.33GB. Estimated download time: 5–10 minutes (varies by network speed). A successful download will show progress like this:
Fetching 4 files: 100%|██████████████████████████████████████████| 4/4 [05:45<00:00, 115.30s/it]Create a `.env` file in the project root directory to store environment-specific variables (e.g., API keys, custom file paths, or model cache directories). Example content:
# Path to the GeneMarkS Perl script (used for gene prediction)
GMS_SCRIPT_PATH="/home/shenmaa/gms2_linux_64/gms2.pl"
# Path to the Java Development Kit (JDK) installation
JAVA_HOME=/data/zhurongpeng-20250919/jdk-25
# OpenAI API key for accessing OpenAI models
OPENAI_API_KEY='sk-xxx'
# Large language model (LLM) used in the RAG pipeline
LLM_MODEL='gpt-4o'
# Embedding model used to generate vector representations of text chunks
EMBEDDING_MODEL='text-embedding-3-small'
# ReRank model used to re-score and reorder retrieved results
RERANK_MODEL='gte-rerank-v2'
# Sampling temperature for LLM responses (lower = more deterministic)
TEMPERATURE=0.3
# Maximum number of tokens allowed in the LLM's response
MAX_TOKENS=2048
# API key for DashScope (Alibaba Cloud) services, used by the ReRank model
DASHSCOPE_API_KEY_ENV='sk-xxx'
# API key for DeepSeek LLM service (alternative or additional LLM provider)
DEEPSEEK_API_KEY='sk-xxx'
# Chunk size (in characters or tokens) for splitting documents in RAG
CHUNK_SIZE=800
# Overlap size between consecutive chunks to preserve context
CHUNK_OVERLAP=120Launch the CASPIA web interface:
python webui.pyThe application will start on http://localhost:7860 (or http://0.0.0.0:7860 for network access). Open this URL in your web browser to access the interface.
- Navigate to the 🤖 CASPIAgent tab
- Type your biological question in natural language
- The agent will process your query using available tools and knowledge bases
- Receive answers with citations and relevant information
Example queries:
- "What is the function of the lacZ gene in E. coli?"
- "Design a plasmid for expressing GFP in yeast"
- "Compare the metabolic pathways of glycolysis in prokaryotes and eukaryotes"
- Navigate to the 🧬 GEMFactory tab
- Upload a genome file (FASTA or GenBank format)
- Configure model parameters (organism type, biomass function, etc.)
- Click "Generate Model" to start automated model construction
- Download the resulting SBML model or view analysis results
- Navigate to the 🔍 CASPIA-RAG tab
- Upload scientific papers (PDF, DOCX, TXT)
- Wait for documents to be processed and indexed
- Ask questions about the uploaded documents
- Receive context-aware answers with source citations
- Navigate to the 📊 Tasks Monitor tab
- View all running and completed tasks
- Check progress, logs, and resource usage
- Download results when tasks complete
Purpose: AI-driven expert agent that orchestrates toolchains for metabolic modeling and strain design.
Core Components:
conversation.py: Multi-turn dialogue and context managementservice.py: Agent planning and task execution logictools/: Encapsulated tool definitions (e.g., gene annotation, model construction, FBA optimization)utils.py: Utility functions for data handling and logging
Supported Backends:
- vLLM-based deployments (Qwen, DeepSeek, OpenAI-compatible models)
- Configurable custom LLM backends
Key Capabilities:
- Natural language interface for complex workflows
- Automated task planning → execution → verification
- Tool-augmented reasoning (e.g., database queries, model optimization)
- Traceable and reproducible report generation
Purpose: End-to-end automated pipeline for constructing parameter-enriched genome-scale metabolic models (ecGEM/etcGEM).
Workflow:
- Genome Annotation: GeneMarkS for ORF prediction → proteome extraction
- Functional Annotation: Protein alignment with Diamond
- Draft Model Construction: CarveMe builds initial stoichiometric GEM
- Parameter Injection: Retrieval from KEGG/BRENDA/BiGG + CASPred predictions (kcat, Topt)
- Validation: Mass-balance, thermodynamic consistency, growth benchmarking
- Optimization:
- Gene-level: FBA, FSEOF, OptKnock strategies
- Protein-level: DMS-based mutation design (MoE PLMs: ProSST, ESM2, ProtSSN, SaProt)
Supported Formats:
- Input: FASTA, GenBank
- Output: SBML, JSON (COBRA standards)
Purpose: High-precision predictive engine for kinetic and thermodynamic parameters missing in GEMs.
Architecture:
- Sequence encoder: ESMC-300M (evolutionary context)
- Structure encoder: Geometric Vector Perceptron (GVP)
- Cross-attention fusion for enzyme–substrate interactions
Key Capabilities:
- Predicts kcat, Topt with uncertainty intervals
- Ensemble learning for confidence estimation
- Continuously updated via wet-lab feedback loop
Integration:
- Called automatically within GEMFactory during parameter completion
- Outputs standardized reports with both values and confidence scores
Purpose: Vision-enhanced Retrieval-Augmented Generation system for scientific literature.
Pipeline:
- Parsing: PDFs → structured Markdown via MinerU
- Vision Enhancement: Image captioning via vision models (charts, figures, tables)
- Chunking & Embedding: Semantic segmentation + vectorization
- Indexing: Stored in ChromaDB for efficient retrieval
- Retrieval: Semantic search + cross-attention re-ranking (Expert Mode)
- Answer Generation: LLM synthesis with citations from both text and images
Features:
- Multi-modal understanding (text + figures + tables)
- Vision-grounded QA with precise references
- Domain-specific optimization for synthetic biology
Purpose: Centralized dashboard for tracking and managing CASPIA workflows.
Features:
- Task queue with scheduling and recovery
- Real-time progress visualization for multi-step jobs
- Resource monitoring (CPU, GPU, memory usage)
- Centralized logging for reproducibility and debugging
- Result aggregation and export for downstream analysis
# Example script for programmatic access (advanced users)
from src.GEMFactory.script.build_model import build_gem
# Build a GEM from genome sequence
model = build_gem(
genome_file="data/ecoli_k12.fasta",
organism_name="Escherichia coli K-12",
gram="negative",
output_format="sbml"
)
# Perform flux balance analysis
from cobra.flux_analysis import flux_variability_analysis
fva_result = flux_variability_analysis(model)
print(fva_result)from src.CASPIA_RAG.agent import RAGAgent
# Initialize RAG agent
agent = RAGAgent(db_path="./src/CASPIA_RAG/db")
# Index documents
agent.index_documents(["paper1.pdf", "paper2.pdf"])
# Query
response = agent.query(
"What are the latest advances in CRISPR base editing?",
top_k=5
)
print(response)from src.CASPIAgent.service import CASPIAgentService
# Initialize agent
agent = CASPIAgentService(model="gpt-4")
# Interactive conversation
response = agent.chat("How can I optimize the production of lycopene in E. coli?")
print(response)SJTU-software-CASPIA/
│
├── webui.py # Main application entry point
├── requirements.txt # Python dependencies
├── requirements_manually.txt # Pytorch & Diamonds dependencies
├── README.md # This file
├── LICENSE # License information
│
├── src/ # Source code modules
│ ├── CASPIAgent/ # Conversational AI agent
│ │ ├── conversation.py
│ │ ├── service.py
│ │ ├── tools.py
│ │ └── utils.py
│ │
│ ├── GEMFactory/ # Metabolic model construction
│ │ ├── data/
│ │ ├── script/
│ │ └── src/
│ │
│ ├── CASPIA_RAG/ # Retrieval-Augmented Generation
│ │ ├── agent.py
│ │ ├── bochaAI.py
│ │ ├── db/
│ │ ├── document/
│ │ ├── image_captioning.py
│ │ ├── load_split_store.py
│ │ ├── prompt.py
│ │ ├── translate.py
│ │ └── util.py
│ │
│ ├── CASPred/ # Prediction modules
│ │
│ └── utils/ # Shared utilities
│
├── tabs/ # Gradio UI tab definitions
│ ├── agent_tab.py
│ ├── gemfactory_tab.py
│ ├── rag_tab.py
│ └── tasks_monitor_tab.py
│
├── static/ # Static assets (images, CSS, etc.)
├── uploads/ # User uploaded files
└── logs/ # Application logs
We welcome contributions from the community! Whether you're fixing bugs, adding new features, or improving documentation, your help is appreciated.
-
Fork the Repository
git clone https://github.com/your-username/SJTU-software-CASPIA.git
-
Create a Feature Branch
git checkout -b feature/your-feature-name
-
Make Your Changes
- Follow PEP 8 style guidelines for Python code
- Add docstrings to all functions and classes
- Include type hints where appropriate
- Write unit tests for new features
-
Commit Your Changes
git add . git commit -m "Add feature: description of your changes"
-
Push to Your Fork
git push origin feature/your-feature-name
-
Open a Pull Request
- Go to the original repository on GitHub
- Click "New Pull Request"
- Provide a clear description of your changes
- Reference any related issues
We welcome contributions from the community! To maintain the reliability and scientific integrity of the CASPIA platform, please follow these guidelines:
-
Code Quality:
- Use clear variable/function names and include docstrings (PEP 257).
- Add type hints wherever possible for better readability and static analysis.
- Ensure deterministic behavior in scientific computations (random seeds, reproducibility checks).
-
Modularity:
- Design components to be reusable across pipelines (e.g., annotation, modeling, RAG).
- Avoid hard-coded paths or organism-specific assumptions.
-
Performance:
- Optimize for efficiency in large-scale AI/ML operations (GPU usage, batching, distributed training).
- Profile heavy tasks (e.g., model inference, database retrieval) before merging.
-
Security:
- Never commit API keys, license files, or other sensitive credentials.
- Be cautious with genome/protein datasets — anonymize or provide public-access examples only.
-
Documentation:
- Update both user-facing docs (README, tutorials) and developer-facing docs (docstrings, comments).
- When adding new features, provide minimal reproducible examples.
If you encounter bugs, inconsistencies, or have feature requests:
- Search first: Check existing issues to avoid duplicates.
- Use templates: Follow the provided GitHub issue templates for bug reports and feature requests.
- Provide details: Include a clear description, minimal reproduction steps, and expected vs. actual behavior.
- System information: Always specify OS, Python version, CUDA version, and GPU model.
- Logs & errors: Paste relevant error messages or stack traces. For long logs, attach as a file or use code blocks.
- Data considerations: If reporting bugs involving biological data, please redact sensitive sequences or genomes and provide synthetic or public test data when possible.
If you use CASPIA in your research, please cite:
@software{caspia2025,
author = {{iGEM SJTU-Software Team}},
title = {CASPIA: Cell-Automated Synthetic Pathway Intelligent Architecture},
year = {2025},
publisher = {iGEM Competition},
url = {https://github.com/shenmaa233/SJTU-software-CASPIA},
version = {v1.0.0-beta},
note = {iGEM 2025 Competition Software Tool}
}2025 iGEM SJTU-Software Team
- Principal Investigators: [To be updated]
- Lead Developers: [To be updated]
We would like to express our gratitude to:
- iGEM Foundation for organizing the International Genetically Engineered Machine competition
- Shanghai Jiao Tong University for institutional support
- Open Source Community for the amazing tools and libraries that made this project possible:
- Our mentors and advisors for their guidance
- Beta testers and early users for valuable feedback
- All contributors who helped improve CASPIA
This project is licensed under the MIT License - see the LICENSE file for details.
This project uses various open-source libraries, each with their own licenses. See LICENSES_THIRD_PARTY.md for details.
- Team Email: adamsthiskywalker@sjtu.edu.cn
- GitHub Issues: Report bugs or request features
- iGEM Wiki: Visit our team wiki
Current Version: 1.0.0-beta
Development Status: Active Development
Last Updated: October 2025
- Core platform architecture
- CASPIAgent module (AI-driven orchestration)
- GEMFactory module (automated parameterized GEMs)
- CASPred module (kinetic/thermodynamic parameter prediction)
- CASPIA-RAG module (vision-enhanced literature QA)
- Tasks Monitor module (workflow tracking & logging)
- API documentation and developer guides
- Docker containerization for reproducible environments
- Cloud deployment support (scalable backend, GPU cluster integration)
- Multi-language UI support (English, Chinese, etc.)
- Dynamic modeling (ODE/DAE integration with GEMs)
- Multi-omics integration (transcriptomics, proteomics, metabolomics)
- Community contribution interface (shared datasets, benchmarks, plugins)
Built with ❤️ by Team SJTU-Software for iGEM 2025